With the looming threat of more drug-resistant pathogens, scientists are looking to untapped resources and ecosystems to find new antimicrobial compounds, Vanessa Zainzinger reports
In past decades, natural active ingredient discovery efforts focused on soil bacteria. Soil-dwelling actinomycetes, for instance, produce almost two-thirds of currently used natural antimicrobial drug compounds.1 Now, however, this intensely explored resource has apparently run dry. Scientists are increasingly isolating already known compounds from soil, prompting them to explore new ecological niches, says Jörn Piel from the Institute of Microbiology at the Swiss Federal Institute of Technology (ETH) in Zurich. Driven by advances in genome sequencing, new natural products with antimicrobial properties are being sought from ever wider sources, from insects and plants to organisms found deep underwater.
Piel has turned his attention to the parts of plants that live above ground, called the phyllosphere. Plant leaves, in particular, are relatively nutrient-poor habitats colonised by a wide diversity of bacteria. ‘Antibiotics might be one of the strategies that bacteria use during interaction and competition for these nutrients,’ Piel explains.In a study2 published in 2018, Piel and colleagues investigated more than 200 species of bacteria living on the leaves of Arabidopsis thaliana, a small weed also known as thale or mouse-ear cress. They chose Arabidopsis because it is widely used as a model organism and has a large library of genetic information backing it up, including decoded genomes of the bacteria that colonise the plant’s leaf surfaces.
Piel’s team found 725 molecular interactions between different strains of bacteria which, in some cases, resulted in preventing bacterial growth. They published a paper on macrobrevin, an active compound with a previously unknown chemical structure, produced by the bacterium Brevibacillus sp. Leaf 182. Of course, finding new antibiotics isn’t quite as straightforward as that; macrobrevin is chemically unstable and not the best candidate for further evaluation, says Piel. But he still has high hopes for the phyllosphere: ‘We currently focus on other, more promising compounds that we have recently discovered. These have novel structures, are highly stable, and active against important pathogens.’
In addition to leaf bacteria, Piel is exploring other poorly studied habitats and organisms, such as sponge microbiomes, other marine bacteria and rhizosphere communities. A particularly fascinating resource is ‘microbial dark matter’: elusive microorganisms from a wide range of taxa that lack cultivated representatives, he says. ‘Here, ours and other labs find evidence for fascinating chemistry that is largely untapped. Method development that allows us to characterise and access the compounds of this rich resource is an important goal.’
|
10,000 |
Amazing ants
Insects are another vast resource that has so far escaped the spotlight. ‘Fifty percent of all species – over a million – are insects. This is potentially a rich source for looking for new antibiotic compounds but it hasn’t been tapped,’ says Clint Penick, an assistant professor at Kennesaw State University, US.
Conventional wisdom in biology dictates that social insects - such as bees, termites, wasps and ants - carry antimicrobial agents. The underlying reasoning is that, living in dense societies, they face a high chance of disease transmission, much like humans living in cities. There has, however, been little concrete evidence for this theory because researchers have been lacking a reliable method for sampling antimicrobial compounds produced by insects at scale.
Penick and co-researchers at the universities of Arizona State, North Carolina State, Campbell, Pennsylvania State and Copenhagen set out to produce an assay to change that.3 The team tested the antimicrobial properties associated with 20 ant species. They used a solvent to remove all of the substances on the surface of each ant’s body, then introduced the resulting solution to a bacterial slurry. Finally, they compared the growth of the bacteria in the slurry with the growth of bacteria in a control group, to see whether an antimicrobial agent was at work.
The study has a plot twist: none of the researchers’ predictions held up. ‘All ants are social so we expected that all of them would produce antimicrobials and that ant species that live in big colonies would produce the strongest antimicrobials. But none of that was true,’ Penick explains.
In fact, half of the ant species Penick tested produced no antimicrobials at all. And the species that produced the strongest antimicrobial compounds – the desert fire ant – lives in some of the smallest colonies.
Penick says these findings open up several avenues for new antibiotics. First, the now tried and tested assay will allow his group to narrow down the groups of ants that do produce interesting antimicrobials. ‘Eventually we’ll be able to pass this research on to chemists who have the tools and knowledge to produce new compounds,’ he says.
|
Derived from a species of marine bacteria that lives deep underwater, marinomycin A is able to defeat infection from both methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant Enterococcus faecium (VREF). |
But his interest is also spiked by those ants that do not produce antimicrobials at all. ‘I wouldn’t be surprised if all the thousands of different species of ants have thousands of different strategies for dealing with pathogens,’ he says.
Currently, Penick’s team is exploring whether some species are culturing microbes that help them to prevent pathogens from taking hold on their bodies. Others might be producing targeted antimicrobials when they are needed and deploying them in a more exact manner rather than blanketing their whole body or nest with it, he says. Yet other species could be forming partnerships with microbes that themselves are producing antimicrobials or outcompeting pathogens, Penick speculates, which would tie in with similar work on insects led by University of Wisconsin-Madison professor Cameron Currie (see Insect Bonanza below).
‘As we dig deeper, I hope we can help expand the idea in humans that antibiotics are not the only solution for dealing with pathogens,’ Penick says.
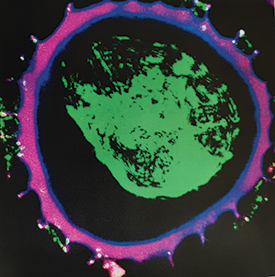
Image: How supercharged deep sea antibiotics (green) appear once encased in their pollen sunscreen shell, in purple.
Image: How supercharged deep sea antibiotics (green) appear once encased in their pollen sunscreen shell, in purple.
Deepwater drugs
In the meantime, the search for new antibiotics continues to dig deeper as well. The deep sea is a relatively unexplored ecosystem that is potentially full of exciting compounds, such as the potent antibiotic agent marinomycin A.
As a potential antibiotic, marinomycin A is as exciting as they come – able to defeat infection from both methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant Enterococcus faecium (VREF). But the compound, which is derived from a species of marine bacteria that lives deep underwater, is so light-sensitive that its delicate polymer structure crumbles in less than two minutes upon exposure to direct UV light, making it useless in a conventional clinical setting.
Earlier in 2019, however, the news that a group of chemists had found a way to turn the chemical into a usable drug was greeted with much excitement.4 The solution to marinomycin A’s light-sensitivity is pollen, according to Andrew Evans from Queen’s University Canada, who hit upon the idea of using pollen shells as a kind of sunscreen for marinomycins. Evans worked on his idea in collaboration with Grahame Mackenzie from the University of Hull, who had developed a technique for replacing the contents of plant spores with active agents, at his company Sporomex; and organic chemist Rebecca Goss from the University of St. Andrews in Scotland, who had already isolated the marinomycins strain.
Encasing marinomycin A in a pollen shell turned out to be easy, Evans says. The solid shells are simply introduced during a fermentation process. Once encased, marinomycin A becomes stable for more than seven hours in direct UV light, which is significantly stronger than regular sunlight.
With these results, a marinomycin A drug suddenly took a leap toward becoming reality. Evans adds that because the spores are naturally occurring, they would not complicate the regulatory approval process for marinomycin A, which can be submitted for clinical trials entirely on its own merits. Nor is this approach limited to one particular type of spore-bearing plant. ‘Different plant species give different pore sizes,’ he says. ‘You can tailor the pore size to the size of molecule, which makes this a fairly versatile strategy.’
The team’s paper was greeted earlier in 2019 with a wave of media attention in the summer. Evans hopes the resulting excitement around the results will help them garner funding to pursue the research further, initially by testing the pollen-encased marinomycin A in animal models.
The positive reactions are even more encouraging because the team conducted its research without any funding to back it up. Evans says funding agencies considered it too risky, and calls on them to be more bold when it comes to research tackling the antibiotics crisis.
‘I think there’s a lot of good ideas that often fall through the cracks because they’re deemed too risky,’ he says. ‘But that’s what science is about: addressing things we don’t know the answer to. And a research area that is really risky could have a huge impact.’
Image (Clint Penick: Foragers of the desert fire ant, Solenopsis xyloni, collecting flower nectar. Ants in this genus produce some of the strongest antimicrobials measured in social insects.
Capable cranberries
While risky research may be rewarded with the discovery of new drugs, bringing them to market takes years, even decades. Antibiotic resistance, however, is a problem now. Researchers at the Canadian Institut National de la Recherche Scientifique (INRS) and McGill University are trying to tackle the issue from a simpler, potentially quicker, angle – with the finding that a compound in cranberries can make pathogenic bacteria more sensitive to existing antibiotics.5
Cranberry proanthocyanidin (cPAC), a polyphenol that gives the fruits their colour, makes the bacterial cell wall more permeable to antibiotics and interferes with the mechanism used by bacteria to pump out the drug. Consequently, the antibiotic penetrates more easily, and bacteria have a harder time getting rid of it. As such, cPAC can decrease the concentration of antibiotic needed to kill bacteria. But it could also help prevent the emergence of resistance, says study co-author, Eric Deziel from INRS. ‘The exciting aspect of adjuvants like cPAC versus antibiotics is they do not impose a survival pressure on the bacteria, because they are not antimicrobials per se,’ he says.
Deziel and his co-researchers tested the cranberry extract, and a range of antibiotics, on bacteria that are responsible for urinary tract infections, pneumonia, and gastroenteritis: Proteus mirabilis, Pseudomonas aeruginosa and Escherichia coli. Bacteria that were exposed to an antibiotic alone developed a tolerance within a few days. But bacteria exposed to a combination of antibiotic and cPAC did not.
It is ‘very likely’ that the extract will have a similar effect on other types of bacteria, Deziel says. cPAC targets general mechanisms used by various classes of bacteria to resist antibiotics and other biocides: selective membrane permeability and multidrug efflux pumps.
The potential results are intriguing. If the researchers can identify the active molecules and pinpoint the exact mode of action, their findings could allow medicine to use reduced doses of antibiotic to treat infections and decrease the likelihood of bacteria becoming more tolerant.
In addition, it might open the possibility of bringing back old antibiotics that are no longer in use because of broad resistance, says Deziel. ‘Indeed, several antibiotics are cheap to produce and low in toxicity but unfortunately no longer in use because most pathogenic bacteria have become resistant. It would be fantastic to reintroduce them.’
|
One interesting source of new antibiotics could be the phyllosphere – the parts of plants that live above ground. Plant leaves, for example, are relatively nutrient-poor habitats, colonised by a wide diversity of bacteria. In 2018, researchers reported finding more than 200 species of bacteria that live on the leaves of Arabidopsis thaliana, also known as mouse-ear cress. |
References
1 S. D. Bentley et al., Nature, 2002, 417, 141.
2 E. J. N. Helfrich et al,. Nature Microbiology, 2018, 3, 909.
3 C. A. Penick et al., Royal Society Open Science, 2018, 5:171332.
4 C. S. Bailey et al., Chemical Science, 2019, 10, 7549.
5 V. B. Maisuria et al., Advanced Science, 2019, 6(15), 1802333.
6 M. G. Chevrette et al., Nature Commun., 2019, 10(1), 1.
|
Insect bonanza In all, the insects provided more than 10,000 microbes to test. In a mammoth effort, the team conducted 50,000 trials to test each microbe’s ability to inhibit the growth of 24 different bacteria and fungi. The work resulted in the discovery of a new antibiotic from a Brazilian fungus-farming ant, which was effective in lab tests against fungi resistant to most other antibiotics and combatted fungal infections without causing toxic side effects in a mouse model. The researchers named it cyphomycin and have submitted a patent. Currie has previously shown that some insect-associated microbes provide their hosts with protection against infections. And, because many insects rely on microbial antibiotics to combat ever-evolving pathogens in their own environment, they have likely selected for antibiotics that can overcome common resistance mechanisms. |